IMPROVEMENT
CollectionContribution


To extend the usage of BioBrick-compatible pBR322-based vector (pSB6A1) we changed the promoter of lac to ldhp and an antibiotic of ampicillin to kanamycin or chloramphenicol to generate two plasmid backbones of pSB6K1 (BBa_K3728009) and pSB6C1 (BBa_K3728010) and a composite part of ldhp-RFP-Tr/pSB6C1 (BBa_K3728013).
In our project, we focus on tackling an issue of food supply and security. Salmonella spp. are pathogens common in foodborne disease outbreaks. Salmonella typhimurium strain LT2 and the pBR322 vector with AmpR are widely used in Salmonella transgenesis studies in the laboratories1. Unexpectedly, the wild type of Salmonella labelled LT2 obtained from the lab of Prof. Cheng-Yang Huang at Chung Shan Medical University survived and grew on LB ampicillin agar plates (now we called it strain CYH). In addition, because of emergence of multidrug-resistant Salmonella2, we are motivated to develop tools for Salmonella spp. including a broad-host-range promoter and antibiotic resistance cassettes based on improving the BioBrick existing part of pBR322-based pSB6A1 (AmpR).
The Salmonella strain CYH was identified by PCR with primer sets based on the sequences in the KEGG database and Park SH’s paper3. Fig. 1 indicated the presence of Salmonella genes and genus-conserved sequences.

Fig. 1. Salmonella Typhimurium LT2 genes and the Salmonella-genus conserved sequence identified on gel electrophoresis with 1 kb marker. Lanes: 1. crr (510 bp), 2. RBS-crr (528 bp), 3. ptsG (1434 bp), 4. RBS-ptsG(1452 bp), 5. STM1128 (1497 bp), 6. RBS- STM1128 (1515 bp), 7. Salmonella-genus conserved gene STM3098 (423 bp)
Then, we tested CYH strain with antibiotics frequently used in iGEM projects. (i.e., ampicillin, chloramphenicol and kanamycin). We found Salmonella strain CYH can grow on LB agar plates supplemented with 100 μg/ml of ampicillin but not with 20 μg/ml of chloramphenicol and 50 μg/ml of kanamycin (Fig. 2).

Fig. 2. Salmonella lab strain tested on LB agar plates with different antibiotics as indicated.
According to the results of antibiotics screening, we built up pSB6K1(BBa_K3728009) and pSB6C1 (BBa_K3728010), where AmpR coding region was replaced by KanR and CmR, respectively (Fig. 3). The plasmid backbones with the intrinsic BBa_J04450 standard parts between Prefix and Suffix were confirmed by growing the transformed E. coli on corresponding LB agar plates and by colony PCR check with primers on Subpart of E1010 (mRFP) and KanR gene or CmR gene (Fig. 4).

Fig. 3. Schematic plasmid maps of BioBrick pBR322-based vectors. pSB6A1 is an existing part. pSB6K1 (BBa_K3728009) and pSB6C1 (BBa_K3728010) are our new parts.
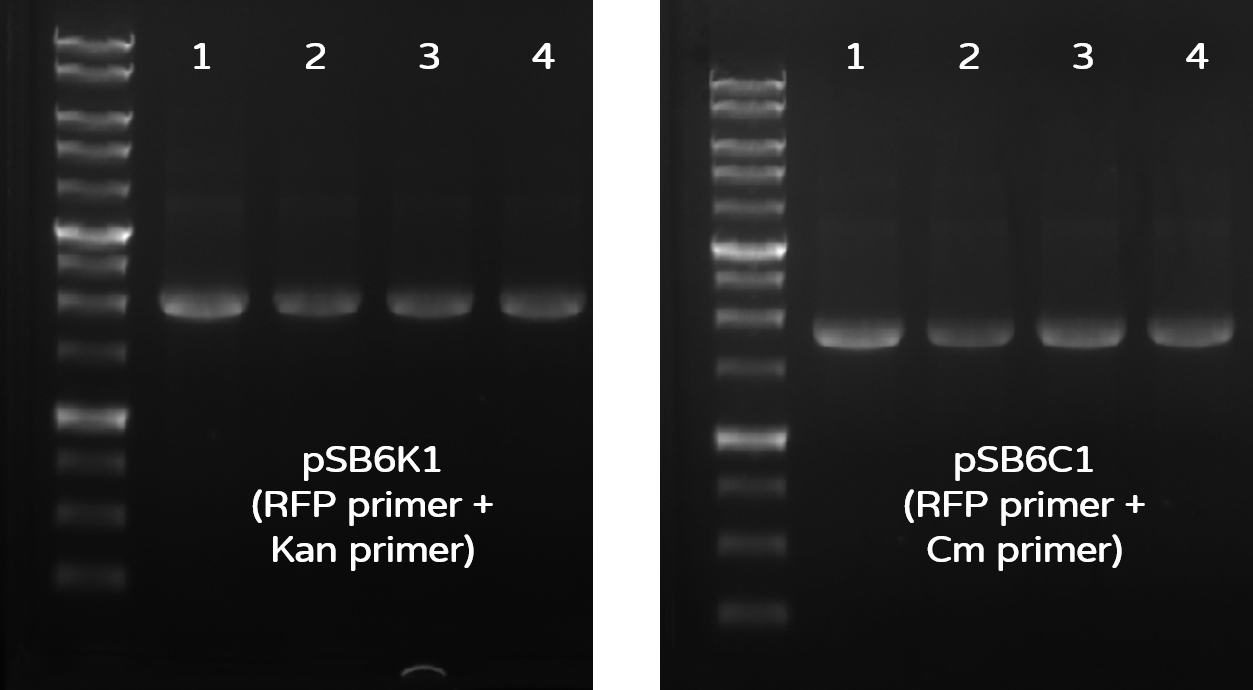
Fig. 4. Colony PCR check results on gel electrophoresis with 1 kb marker. Left gel: PCR check of 4 colonies for pSB6K1 with RFP-R + KanR-R to generate 1950-bp bands. Right gel: PCR check of 4 colonies for pSB6C1 with RFP-R + CmR-R to generate 1806-bp bands.
ldhp ( BBa_K3376000) is a strong and constitutive promoter created by us in iGEM 2020. Its activities were characterized in in vitro transcription-translation (TXTL) assay and in E. coli, Salmonella spp. and Gram-positive bacteria such as S. mutans. Moreover, pBR322-based vector is widely used to create shuttle vectors between Gram negative and positive bacteria4. Therefore, we further extended the usage of the vector by addition of the broad-host-range promoter of ldhp to replace the lac promoter. The resulting ldhp-RFP-Tr/pSB6C1 (BBa_K3376013) was confirmed by restriction enzyme check (Fig. 5).

Fig. 5. Restriction enzyme check of ldhp-RFP-Tr/pSB6C1 with EcoRI and PstI. The plasmids were prepared from 3 colonies of overnight culture of the transformed E. coli. Gel showed the 3817-bp and 1092-bp bands by restriction enzyme cuts.
Finally, the ldhp-RFP-Tr/pSB6C1 is able to transform Salmonella by electroporation at a single pulse of 12.5 kV/cm (2.5kV, 200Ω, 25μF) 1 to become chloramphenicol resistant and red colonies on a LB agar plate (Fig. 6), whereas the existing part of pSB6A1 can not be used in an ampicillin-resistant Salmonella strain.

Fig. 6. Salmonella was transformed with the vectors of pSB6C1 (CmR) or pSB6A1 (AmpR) as a control. The result showed red colonies formed on a Cm LB agar plate with ldhp-RFP-Tr/pSB6C1 and no colonies with pSB6A1 vector.
Go to check our documentation on the existing Part:pSB6A1 (pBR322 converted to BioBrick vector) and our new Part:BBa_K3728013 (ldhp-RFP-Tr/pSB6C1)

DOWNLOAD HERE👇👇👇
1. O'Callaghan D, Charbit A. High efficiency transformation of Salmonella typhimurium and Salmonella typhi by electroporation. Mol Gen Genet. 1990 Aug;223(1):156-8. doi: 10.1007/BF00315809.
2. V T Nair D, Venkitanarayanan K, Kollanoor Johny A. Antibiotic-Resistant Salmonella in the Food Supply and the Potential Role of Antibiotic Alternatives for Control. Foods. 2018 Oct 11;7(10):167. doi: 10.3390/foods7100167.
3. Park SH, Ricke SC. Development of multiplex PCR assay for simultaneous detection of Salmonella genus, Salmonella subspecies I, Salm. Enteritidis, Salm. Heidelberg and Salm. Typhimurium. J Appl Microbiol. 2015 Jan;118(1):152-60. doi: 10.1111/jam.12678.
4. P. Trieu-Cuot, C. Carlier, P. Martin, P. Courvalin, Plasmid transfer by conjugation from Escherichia coli to Gram-positive bacteria, FEMS Microbiology Letters. 1987 Dec;48(1-2):289-294. doi: 10.1111/j.1574-6968.1987.tb02558.x.