
Overview
In our project, the superfolder green fluorescent protein (sfGFP) allows better quantification of promoter strength and sensitivity.[1,2]. In the oxidative stress sensing system biobrick, we improved the biobrick BBa_K2610031 from the 2019 iGEM Leiden team by changing GFP (BBa_E0040)[2]. into sfGFP(BBa_I746916), which is hypothesized to have a higher expression level than GFP [Fig. 1]. We also added transcription activator SoxR to our biobrick for increased function of the oxidative stress sensing system[3].
Results
Disk assay
Disk assay was used to check the effect of each inducer. The concentration of the different inducers we used are listed below:
| Inducer | Volume per disk |
|---|---|
| H2O2s (30%) | 10μl |
| H2O2w (3%) | 10μl |
| MD (menadione)(10mM) | 10μl |
| DMSO (solvent for MD) | 10μl |
| PQ (paraquat)(1mM) | 10μl |
| MQ (solvent for PQ) | 10μl |
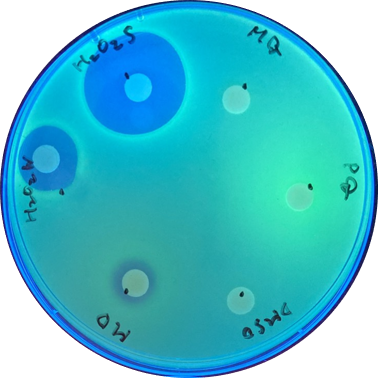
With sfGFP, the induced system result can be checked under UV light, making it easier to tell the difference between each inducer. [Fig 2.]
Finally, to confirm the protein activity of TAL and TyrP, we performed a functional test using n-octanol extraction method, which was previously proposed by iGEM_Uppsala 2013 and has been verified by HPLC[3]. The p-Coumaric acid concentration was measured through the absorbance value at 310 nm wavelength under Nanodrop UV-Vis wavelength.
We compared the TAL constructs containing the native and B0034 ribosome binding sites, (BBa_K2997009 and BBa_K2997010) to determine if p-Coumaric Acid production is improved by changing the ribosome binding sites. From the results seen in Fig. 4, BBa_K2997010 is able to produce a higher amount of p-Coumaric acid. Hence, we are able to prove that by changing the RBS (from Native to B0034), the conversion of tyrosine into p-Coumaric acid can increase by 1.73-fold. Therefore, we have shown that we have improved a previous BioBrick.
For more information, please visit our Results page.
References
- https://2007.igem.org/wiki/index.php/Edinburgh/Team
- Chen, H., Bjerknes, M., Kumar, R., & Jay, E. (1994). Determination of the optimal aligned spacing between the Shine – Dalgarno sequence and the translation initiation codon of Escherichia coli mRNAs. Nucleic Acids Research, 22(23), 4953–4957.
- https://2013.igem.org/Team:Uppsala



