
Overview
Toggle macrophage is a smart CAR macrophage designed to phagocytose SARS-CoV-2 and switch its regulatory functions on inflammation automatically according to the IL-6 levels. We endowed Toggle Macrophage with a synthetic system consisting of:
1.IL-6 sensor 1 and IL-6 sensor 2: The IL-6 sensors are capable of sensing the IL-6 levels and releasing transcriptional factors to control the downstream gene expression.
2.CARγ: Promote phagocytosis and inflammation
3.CARMERTK: Promote phagocytosis and inhibit inflammation
4.Kill switch: Induce suicide of Toggle Macrophage. Please scroll down to see more details in our works.
Circuit Design for Toggle Macrophage:
Toggle macrophage is a chimeric antigen receptor (CAR)-macrophage engineered with IL-6 sensor. We chose macrophages (M0) as the chassis because:
1.Macrophages participate in the innate immune response and play a critical role in the body’s defense against viral infections. They are powerful activators for triggering adaptive immune responses and killing viruses (Meidaninikjeh et al. 2021).
2. Naturally, they can transform into M1 type and M2 type, which is pro-inflammatory and anti-inflammatory, respectively. This characteristic of macrophages corresponds to our ideal therapy which eradicates the viruses and balances the immune response at the same time.
3.Delayed IFN-I release is associated with the inactivation of macrophages, which later causes delayed viral clearance and increased proinflammatory cytokine release.
4.There is no reported evidence for the presence of infectious viruses or toxic products in THP-1 cells, making this cell line relatively easy and safe to use (Chanput, Mes, and Wichers 2014).
5.Recently, researchers have constructed CAR-macrophages, which exerts their efficacy of killing the cancer cells. CAR-macrophages were endowed with enhanced recognizing ability and phagocytosis. Therefore, we considered that CAR-macrophages would be a strong tool to treat SARS-CoV-2 infection.
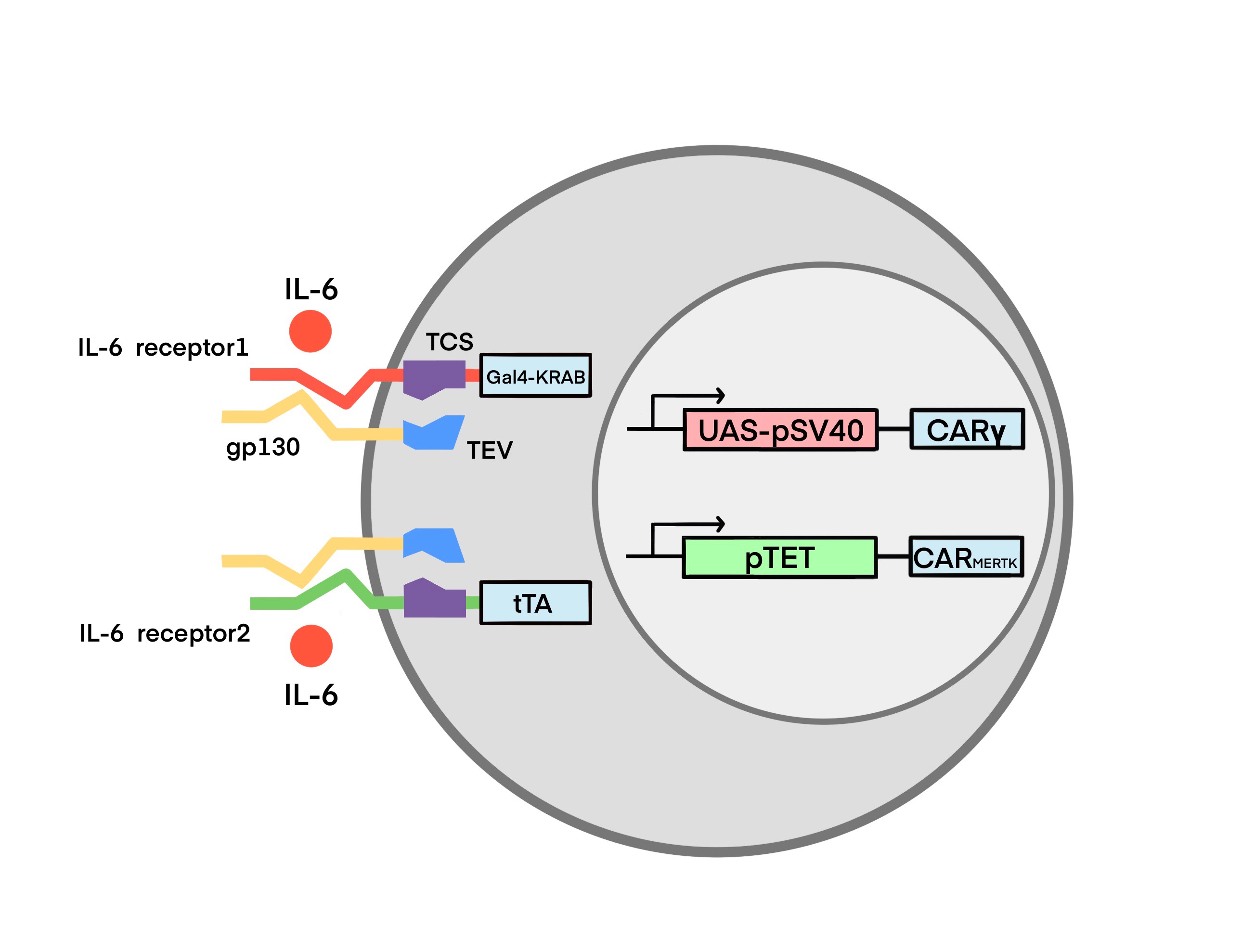
To promote the recognizing ability and phagocytosis of macrophage, we engineered macrophages with CAR. By designing the intracellular domain of the CAR, we obtained two CARs: CARγ and CARMERTK. We integrated the two CARs and IL-6 sensors into macrophages and achieved a smart CAR-macrophage, named Toggle Macrophage. About the test of intergration, see in Proof of Concept. The whole circuit regulates the functions of Toggle Macrophage by switching the CAR expression based on the IL-6 concentration in the microenvironment. When the IL-6 concentration is low, the CARγ-macrophage (Mγ) dominate, which clear the viruses and enhance the release of proinflammatory cytokine. In contrary, when IL-6 concentration is high, the CARMERTK-macrophage (Ms) dominate, which eradicate the viruses and suppress the inflammatory response.
Below, we describe how Toggle Macrophage works in detail, including its dynamic response to inflammation.

As described above, neither IL-6 receptor is triggered when the IL-6 concentration in the body does not reach a threshold, and the transcription factors Gal4-KRAB (IL-6 receptor 1) and tTA (IL-6 receptor 2) are not released. The following effects occur.
a. IL-6 receptor 1 does not trigger: initiating CARγ transcription
1. pSV40 promoter activates transcription of the gene aSscFV-Fcγ.
2. CARγ is abundantly expressed on the cell membrane.
b. IL-6 receptor 2 does not trigger: repressing CARMERTK transcription
1. PminCMV is highly inefficient and fails to initiate transcription.
2. CARMERTK is barely expressed on the cell membrane
As can be seen, the inactivation of IL-6 receptor 1 and 2 results in enhancing CARγ expression and suppressing CARMERTK expression, respectively. In general, these effects promote the conversion of toggle macrophages into Mγ. The proportion of Mγ in the macrophage cluster gradually rises and results in phagocytosing the viruses and enhancing the immune function of the patient.

As described above, when the IL-6 concentration within the environment reaches a threshold, both IL-6 receptors are triggered and the transcription factors Gal4-KRAB (IL-6 receptor 1) and tTA (IL-6 receptor 2) are released. The following effects are produced.
c. IL-6 receptor 1 triggers: suppressing CARγ transcription
1. Gal4 binds to UAS.
2. KRAB represses the expression of the downstream gene aSscFV-Fcγ.
d. IL-6 receptor 2 triggers: promoting CARMERTK transcription
1.tetR binds to tetO.
2.VP64 binds to the corresponding cis-acting elements.
3.transcription of the gene aSscFV-MERTK is then initiated by PminCMV downstream of TRE.
The activation of IL-6 receptor 1 and 2 results in repressing CARγ expression and promoting CARMERTK expression, respectively. In general, these effects promote the conversion of toggle macrophages into Ms. The proportion of Ms in the macrophage cluster gradually rises and results in phagocytosing the viruses and suppressing the cytokine storm.

Ms then causes a gradual decrease in IL-6 concentration, the transcription of CARγ gene restarts, while the transcription of CARMERTK gene gradually stops. Correspondingly, the Ms gradually apoptosis, while the Mγ gradually proliferate, and finally the macrophage cluster reassumes the function of Mγ again, forming an effective negative feedback loop regulatory mechanism. Ultimately, we hope to reach a ‘cytokinostasis’, keeping the IL-6 concentration within a safe range through the intelligent conversion of macrophages.

SARS-CoV-2 causes various symptoms in patients that develop dynamically in accordance with the course of COVID-19, necessitating the use of a smart therapy like Toggle Macrophage. From previous literature review, we learned that SARS-COV-2 exhibited immune escape in the early stage of the infection. The lack of the timely immune response augments the risk of developing into severe cases(Chau et al. 2021). Therefore, it’s highly important to increase the expression of the cytokines in the early stage so that more immune cells can be recruited to defend against viruses. However, unlike early-stage patients, those who were in severe cases may be susceptible to the cytokine release syndrome (CRS). The high levels of peripheral proinflammatory cytokines can be devastating to the host. Inspired by these, we decided to design a novel CAR-macrophage which can respond intelligently and dynamically to varying degrees of inflammation and stages of infection. For mild or asymptomatic patient, the Toggle Macrophage will transform into the Mγ. It can eradicate the viruses by redirecting the S protein and strengthen the immune response by secreting proinflammatory cytokines. For patients experiencing CRS, Ms enable to ameliorate their conditions. In the presence of excessive proinflammatory cytokines, the IL-6 sensor will activate. It sends a signal to inhibit CARγ expression and a signal to activate CARMERTK expression. Eventually, the macrophage cluster converts to the state of Ms which limits the systemic inflammation when killing the viruses.
To provide a therapeutic effect, Toggle Macrophage relies on five modules: CARγ, CARMERTK, IL-6 sensor1 , IL-6 sensor2, and a kill switch. The individual modules are described below.
Module 1 Phenotypic CAR Receptors:
The core of our design is to find the dynamic balance between CARγ and CARMERTK, the output of the two genetic circuits. In the early stage, when the levels of IL-6 is below the detection threshold of IL-6 sensor, CARγ will be the main force to combat the virus through increasing the expression of cytokines and impairing immune escape. However, CARMERTK will exceed CARγ in number when IL-6 sensor detects high levels of IL-6, performing a strong phagocytosis of SARS-COV-2 and keeping immunologically silent as much as possible.
1. The extracellular domain: a single chain variable fragment (ScFv) which specifically recognizes the spike protein of SARS-CoV-2.
2. The transmembrane domain: CD8 protein allowing signal transduction.
3. The intracellular domain: This domain consists of different signal motifs which stem from the intracellular structure of various natural receptors on the surface of immune cells. These signal motifs can trigger robust phagocytosis.
4. Myctag is a fluorescent protein gene for subsequent detection of CAR macrophage transduction.

According to research,MERTK is a tyrosine kinase which belongs to TAM family of RTKs. The activated TAM receptors can inhibit the TLR and type I IFN signaling pathways and thus may avoid further upregulation of proinflammatory cytokine levels(Todt, Hu, and Curtis 2004). In previous study, Megf10, and FcRγ robustly triggered engulfment of CAR-macrophages by being phosphorylated by Src family kinases, while CD3ζ by recruiting syk kinase(Morrissey et al. 2018).
We further conducted several experiments to screen out the ideal CAR (see in Engineering Success). Based on that we finally chose CARγ and CARMERTK as our utilized CAR.
Module 2 Cytokine Sensor:
Recent studies have shown that overproduction of several cytokines is associated with poor prognosis of COVID-19(Cao 2020). Yun Ling et al. found that plasmatic concentrations of IL-5, IL-6, and IL-8 were significantly elevated. These cytokines are predominant in cytokine storm in dead cases(Liu et al. 2021). Clinically, IL-6 levels are considered to be a predictor of severity and prognosis in COVID-19(Tang et al. 2021). Based on these facts, we selected IL-6 as an indicator of inflammation and designed two IL-6 receptors to control the expression of CAR. (referred to as IL-6 receptor 1, IL-6 receptor 2, respectively)
The two IL-6 receptors are mostly identical in design, the only difference is that IL-6 receptor 1 is downstream of transcription repression factor, GAL4-KRAB fusion protein, while IL-6 receptor 2 is downstream of transcription activation factor, tTA. In addition, IL-6 receptor 1 regulates CARγ expression, while IL-6 receptor 2 regulates CAR METK expression.

The IL-6 Receptor Complex
The IL-6 receptor complex consists of the IL-6-specific alpha receptor, IL-6R α (gp80, CD126) and the signal transduction subunit glycoprotein130 (gp130, CD130). Notably, IL-6Rα chains exist in soluble or membrane form. Binding of IL-6 to membrane IL-6 receptor α chains generates anti-inflammatory classical signals, whereas binding to soluble IL-6 receptor α chains generates pro-inflammatory signals.
In the classical signaling pathway, IL-6 first binds with low affinity to the specific membrane receptor IL-6R (CD126), and the IL-6-IL-6R complex binds with high affinity to the signaling receptor component glycoprotein 130 (gp130), which is homodimerized into a hexamer(Reeh et al. 2019) (containing two each of IL-6, IL-6R, and GP130).

Based on the high affinity binding of IL-6 to IL-6R and gp130 expressed by monocytes(Hunter and Jones 2015). We designed the IL-6 receptor with two parts. The first part consists of a membrane-type gp130 with tobacco etch virus protease (TEVp). TEVp is a highly specific cysteine protease in TEV. A number of TEVp mutants with different rate of cleavage, stability and specificity have been reported. Similarly, a panel of different TEV cleavage sites(TCS), derived from the canonical ENLYFQ-G/S site, has been established. TEVp is highly valued for its good transduction efficiency and specificity. To our knowledge, it was never reported that TEVp would cleave fusion proteins out of target. Moreover, TEVp harbors several advantages in biotechnological applications, such as simple production process, low cost, and utility of open source vectors and mutants, making it suitable for different in vitro and in vivo applications(Cesaratto, Burrone, and Petris 2016).

The second part consists of the extracellular domain of membrane IL-6R, TEV cleavage site (TCS) and transcription factors. The transcription factors are the transcriptional repressor GAL4-KRAB (IL-6 receptor 1) and the transcriptional activator tTA (IL-6 receptor 2), respectively. Specific information about GAL4-KRAB and tTA will be described in the Module 3.


When the IL-6 concentration in the environment reaches a certain threshold, IL-6 receptors are activated as follows: (Ps: For both receptors, the triggering process is the same)
1.IL-6 binds to membrane IL-6R (mIL-6R) with low affinity.
2. IL-6-IL-6R complex binds to gp130 extracellularly.
3.TEV protease downstream of gp130 is activated and hydrolyzes the TCS.
4. The transcription factor Gal4-KRAB (IL-6 receptor 1)/tTA (IL-6 receptor 2) is released.
Module 3 Linked Gene Circuits:

GAL4-KRAB:
This part is a mammalian synthetic transcription factors based on Gal4 DNA binding domain (DBD)(Pengue et al. 1994) and KRAB transcription repression domain. Gal4-KRAB containing three core domains from N-terminal to C-terminal: GAL4 DNA binding domain, nuclear location sequence (NLS) and KRAB transcription repression domain(Morsut et al. 2016). And a (G4S) linker was added between DBD and NLS for providing region flexibility(Witzgall et al. 1994; Chen, Zaro, and Shen 2013). GAL4 DBD is capable of binding to specific DNA sequences and KRAB repressing the expression of downstream gene, so that we used Gal4-KRAB as a transcription repression factor to inhibit the activation of downstream synthetic promoter UAS-pSV40.
1.KRAB(Kruppel-associated box): KRAB is the N-terminal reserve area of Kruppel-associated protein, Kid-1, which can reversibly inhibit the gene expression by forming heterochromatin via recruiting various histone mediation factors(Wang et al. 1997). The KRAB protein has been demonstrated to be capable of inhibiting all promoters within at least 3 kB(Deuschle, Meyer, and Thiesen 1995). Strong transcriptional repression was observed when the KRAB domain was bound both at near or kilobase distances from the start site of transcription.
2.Gal4: Gal4 is a yeast transcriptional activator consisting of 881 amino acid. The DNA binding activity of Gal4 is located in the 74 amino acids in the N terminus (Keegan et al. 1986). The transcriptional activation function of Gal4 is mapped in two regions (residues 148–196 and 768–881) (Ma & Ptashne 1987). Gal4 binds to its specific recognition sequence UAS (upstream activating sequence) and activates transcription of target genes. It has been demonstrated that the Gal4-UAS system can operate not only in yeast but also in various animal cells(Fischer et al. 1988).
UAS-pSV40-CARγ:
1.GAL4 upstream activation sequence (UAS): In cells where the Gal4 gene is expressed, the Gal4 protein targets the UAS sequence(Guarente, Yocum, and Gifford 1982), thus driving expression of any open reading frame immediately downstream of the UAS. In principle, this allows for any gene (linked to UAS) to be expressed in any pattern, so long as there is a known enhancer driving Gal4(Scott 2009; Wang et al. 1997).
2.SV40 promoter (pSV40): the pSV40 is expressed well in the fission yeast S. pombe, and it initiates transcription at the same site as in mammalian cells. In mammalian cells, SV40 early gene transcription is predominantly initiated from a set of closely adjacent sites 24-27 bp downstream of the TATA sequence(Benoist and Chambon 1981)
3.aSscFV-Fcγ(CAR γ): see in Module 1.

tTA (tetR-VP64):
tTA contains three core domains from N-terminal to C-terminal: DNA-binding transcriptional repressor (tetR), NLS and VP64 transcription activation domain. And a linker was added between DBD and NLS for providing region flexibility. tetR DBD is capable of binding to specific DNA sequences and VP64 activating the expression of downstream gene, so that we used tetR-VP64 as transcription activation factor to activate our downstream synthetic cis-acting elements and promotor PminCMV.
1.Tetracycline repressor (tetR): tetR can negatively regulate the transcription of resistance-mediating genes in E.coli. In the presence of the antibiotics tetracycline or doxycycline (Dox), tetR does not bind to its operators (tetO) located within the promoter region of the operon and allows transcription(Gossen and Bujard 1992). In the absence of the antibiotics, tetR binds to tetO and thus represses transcription. Herein, tetR acts as the DBD of tTA, which can specifically bind to TRE (fused with seven tetO)(Gossen and Bujard 1992).
2.VP64: VP64 is a fusion of four VP16, which is more potent than VP16. VP16 is an activating factor that primes transcription from the five virally encoded immediate early (IE) genes in HSV-1(Triezenberg, Kingsbury, and McKnight 1988). The natural tetR is a transcriptional repressor protein in E. coli. Fusing VP64 to tetR can transform tetR can be modified into a transcriptional activator protein available in eukaryotes(Gossen and Bujard 1992). Herein, VP64 acts as the transcription activation domain (AD) of tTA.
pTET (TRE-PminCMV)-CARMERTK:
1.TRE: TRE is the fusion of seven tetracycline operators (tetOs). tetO can be bound with tetR. When bound to tetOs placed upstream of minimal promoters, tTA efficiently activates transcription from such promoters(Gossen and Bujard 1992).
2.PminCMV: PminCMV downstream of TRE is a CMV promoter lacking an enhancer termed minimal CMV promoter, which is inefficient in initiating transcription and can be approximated as being unable to initiate transcription without the combination of tetR and tetO(Gossen and Bujard 1992).
3.aSscFV-MERTK (CARMERTK): see in Module 1.

Illustration of Tet-off system. pTET (BBa_K1061013) consists of TRE and PminCMV. Tandem TetO sequences are positioned upstream of the PminCMV followed by cDNA of gene of interest. Here, a chimeric protein consisting of TetR and VP64 (tTA) is converted into a transcriptional activator, and the expression plasmid is transfected together with the operator plasmid. Thus, culturing cells with Dox switches off the exogenous gene expression, while removing Dox switches it on(Kallunki et al. 2019).
Kill Switch
To ensure that our Toggle Macrophage is safe within the body, we incorporate a chemically-inducible kill switch. This way, the Toggle Macrophage can rapidly be eliminated in vivo. The AP1903 is an excellent candidate for this inducible kill switch (Clackson, et al., 1998), as AP1903 has no other biologic effects in vivo(Straathof, K. C., et al.,2016). Remodeled dimers such as AP1903 are ideal reagents for controlling the activities of cells that have been modified by gene therapy procedures, without interference from endogenous FKBP(Clackson, et al., 1998).
Caspases are very important regulators of apoptosis induced by apoptosis stimuli (Stennicke and Salvesen, 1998). iCasp9 can be activated by AP1903 that has proven safe at the required dose for optimum deletional effect(Iuliucci, J. D., et al.,2001). Caspase9 will subsequently activate downstream effector caspases, such as caspase3, and ultimately induce apoptosis.

Future directions, Novel CAR Receptors, and Alternative Circuits
Our Toggle Macrophage enables to sense the concentration of IL-6 and automatically responds to the developing inflammation in the host. Consequently, Toggle Macrophage control the proinflammatory cytokine level within a specific range. Based on our design, we hypothesize that the system used by Toggle Macrophage as well as Toggle Macrophage itself can be applied to the treatment of other cytokine-related diseases. For example, CRS is one of the main challenges that hinders the wider application of CAR-T therapy in cancers. Notably, a study showed that macrophages produced core cytokines in CRS and played a key role in the CAR T cell associated tumor microenvironment CRS pathogenesis(Hao et al. 2020; Giavridis et al. 2018). Thus, it is possible to engineer the myeloid-derived macrophages with our system, in order to help combat against cancer cells and ameliorate CRS.
To cope with the toxicity of CAR-T therapy, researchers have developed ‘On/off switches’ system to control CAR expression or activity. For instance, Wu et al. constructed a small molecule–gated chimeric receptor that has a split architecture. The specific small molecules act as a bridge between the two split intracellular domains and help form a dimer, thus allowing further signal transduction(Wu et al. 2015). Interestingly, by replacing the chemical dimerization domains with light-inducible dimerization modules, the CAR can be controlled by light. This concept provides us a convenient and sensitive method to regulate the CAR activity(Tan et al. 2017). To enhance the specificity of CAR therapy, researchers introduced another co-stimulatory/co-inhibitory receptor to target multiple antigens. Another strategy is to employ logic gating expression systems, such like synthetic Notch (synNotch) receptors(Rafiq, Hackett, and Brentjens 2020). To overcome the heterogeneity of tumor cells, researchers constructed a bispecific T-cell engager (BiTE) to target two various antigens. BiTE also enables to activate bystander T cells.(Choi et al. 2019). Similar rationale has also been applied to create other novel T cells, such as tandem CAR(Rafiq, Hackett, and Brentjens 2020).
The IL-6 sensor in Toggle Macrophage is designed based on the Tango/MESA synthetic receptor(Schwarz et al. 2017; Edelstein et al. 2020). Roybal et al. constructed a synthetic Notch (synNotch) receptor in T cells. SynNotch receptors contain the core regulatory domain from the cell-cell signaling receptor Notch, but have synthetic extracellular recognition domains (e.g., single-chain antibodies) and synthetic intracellular transcriptional domains. The researchers substituted the target genes downstream of transcriptional factors to impact on the cytokine profile and differentiation of T cells. The engineered T cells can produce specific antibodies or other therapeutics with the regulation of SynNotch receptors(Roybal et al. 2016). Based on these, we deemed that SynNotch is an alternative for IL-6 sensor.
Reference
Benoist, C., and P. Chambon. 1981. 'INVIVO SEQUENCE REQUIREMENTS OF THE SV40 EARLY PROMOTER REGION', Nature, 290: 304-10.
Cao, Xuetao. 2020. 'COVID-19: immunopathology and its implications for therapy', Nature Reviews Immunology, 20: 269-70.
Cesaratto, Francesca, Oscar R. Burrone, and Gianluca Petris. 2016. 'Tobacco Etch Virus protease: A shortcut across biotechnologies', Journal of Biotechnology, 231: 239-49.
Chau, A. S., A. G. Weber, N. I. Maria, S. Narain, A. Liu, N. Hajizadeh, P. Malhotra, O. Bloom, G. Marder, and B. Kaplan. 2021. 'The Longitudinal Immune Response to Coronavirus Disease 2019: Chasing the Cytokine Storm', Arthritis Rheumatol, 73: 23-35.
Chen, Xiaoying, Jennica L. Zaro, and Wei-Chiang Shen. 2013. 'Fusion protein linkers: Property, design and functionality', Advanced Drug Delivery Reviews, 65: 1357-69.
Choi, B. D., X. L. Yu, A. P. Castano, A. A. Bouffard, A. Schmidts, R. C. Larson, S. R. Bailey, A. C. Boroughs, M. J. Frigault, M. B. Leick, I. Scarfo, C. L. Cetrulo, S. Demehri, B. V. Nahed, D. P. Cahill, H. Wakimoto, W. T. Curry, B. S. Carter, and M. V. Maus. 2019. 'CAR-T cells secreting BiTEs circumvent antigen escape without detectable toxicity', Nature Biotechnology, 37: 1049-+.
Deuschle, U, W K Meyer, and H J Thiesen. 1995. 'Tetracycline-reversible silencing of eukaryotic promoters', Molecular and Cellular Biology, 15: 1907-14.
Edelstein, H. I., P. S. Donahue, J. J. Muldoon, A. K. Kang, T. B. Dolberg, L. M. Battaglia, E. R. Allchin, M. Hong, and J. N. Leonard. 2020. 'Elucidation and refinement of synthetic receptor mechanisms', Synth Biol (Oxf), 5: ysaa017.
Fischer, Janice A., Edward Giniger, Tom Maniatis, and Mark Ptashne. 1988. 'GAL4 activates transcription in Drosophila', Nature, 332: 853-56.
Giavridis, Theodoros, Sjoukje J. C. van der Stegen, Justin Eyquem, Mohamad Hamieh, Alessandra Piersigilli, and Michel Sadelain. 2018. 'CAR T cell-induced cytokine release syndrome is mediated by macrophages and abated by IL-1 blockade', Nature Medicine, 24: 731-+.
Gossen, M., and H. Bujard. 1992. 'TIGHT CONTROL OF GENE-EXPRESSION IN MAMMALIAN-CELLS BY TETRACYCLINE-RESPONSIVE PROMOTERS', Proceedings of the National Academy of Sciences of the United States of America, 89: 5547-51.
Guarente, L., R. R. Yocum, and P. Gifford. 1982. 'A GAL10-CYC1 HYBRID YEAST PROMOTER IDENTIFIES THE GAL4 REGULATORY REGION AS AN UPSTREAM SITE', Proceedings of the National Academy of Sciences of the United States of America-Biological Sciences, 79: 7410-14.
Hao, Zhaonian, Ruyuan Li, Li Meng, Zhiqiang Han, and Zhenya Hong. 2020. 'Macrophage, the potential key mediator in CAR-T related CRS', Experimental Hematology & Oncology, 9.
Hunter, Christopher A., and Simon A. Jones. 2015. 'IL-6 as a keystone cytokine in health and disease', Nature Immunology, 16: 448-57.
Kallunki, Tuula, Marin Barisic, Marja Jäättelä, and Bin Liu. 2019. 'How to Choose the Right Inducible Gene Expression System for Mammalian Studies?', Cells, 8: 796.
Liu, Yili, Daihong Chen, Junjie Hou, Haicong Li, Dan Cao, Mingquan Guo, Yun Ling, Menglu Gao, Yi Zhou, Yanmin Wan, and Zhaoqin Zhu. 2021. 'An inter-correlated cytokine network identified at the center of cytokine storm predicted COVID-19 prognosis', Cytokine, 138.
Morrissey, M. A., A. P. Williamson, A. M. Steinbach, E. W. Roberts, N. Kern, M. B. Headley, and R. D. Vale. 2018. 'Chimeric antigen receptors that trigger phagocytosis', Elife, 7.
Morsut, Leonardo, Kole T. Roybal, Xin Xiong, Russell M. Gordley, Scott M. Coyle, Matthew Thomson, and Wendell A. Lim. 2016. 'Engineering Customized Cell Sensing and Response Behaviors Using Synthetic Notch Receptors', Cell, 164: 780-91.
Pengue, Gina, Viola Calabró, Paola Cannada Bartoli, Alfredo Pagliuca, and Luigi Lania. 1994. 'Repression of transcriptional activity at a distance by the evolutionary conserved KRAB domain present in a subfamily of zinc finger proteins', Nucleic Acids Research, 22: 2908-14.
Rafiq, S., C. S. Hackett, and R. J. Brentjens. 2020. 'Engineering strategies to overcome the current roadblocks in CAR T cell therapy', Nature Reviews Clinical Oncology, 17: 147-67.
Reeh, H., N. Rudolph, U. Billing, H. Christen, S. Streif, E. Bullinger, M. Schliemann-Bullinger, R. Findeisen, F. Schaper, H. J. Huber, and A. Dittrich. 2019. 'Response to IL-6 trans- and IL-6 classic signalling is determined by the ratio of the IL-6 receptor alpha to gp130 expression: fusing experimental insights and dynamic modelling', Cell Commun Signal, 17: 46.
Roybal, K. T., J. Z. Williams, L. Morsut, L. J. Rupp, I. Kolinko, J. H. Choe, W. J. Walker, K. A. McNally, and W. A. Lim. 2016. 'Engineering T Cells with Customized Therapeutic Response Programs Using Synthetic Notch Receptors', Cell, 167: 419-32 e16.
Schwarz, K. A., N. M. Daringer, T. B. Dolberg, and J. N. Leonard. 2017. 'Rewiring human cellular input-output using modular extracellular sensors', Nature Chemical Biology, 13: 202-09.
Scott, Ethan K. 2009. 'The Gal4/UAS toolbox in zebrafish: new approaches for defining behavioral circuits', Journal of Neurochemistry, 110: 441-56.
Tan, P., L. He, G. Han, and Y. B. Zhou. 2017. 'Optogenetic Immunomodulation: Shedding Light on Antitumor Immunity', Trends in Biotechnology, 35: 215-26.
Tang, Yueting, Jiayu Sun, Huaqin Pan, Fen Yao, Yumeng Yuan, Mi Zeng, Guangming Ye, Gui Yang, Bokun Zheng, Junli Fan, Yunbao Pan, Ziwu Zhao, Shuang Guo, Yinjuan Liu, Fanlu Liao, Yongwei Duan, Xiaoyang Jiao, and Yirong Li. 2021. 'Aberrant cytokine expression in COVID-19 patients: Associations between cytokines and disease severity', Cytokine, 143.
Todt, J. C., B. Hu, and J. L. Curtis. 2004. 'The receptor tyrosine kinaseMERTK activates phospholipase C gamma 2 during recognition of apoptotic thymocytes by murine macrophages', Journal of Leukocyte Biology, 75: 705-13.
Triezenberg, S. J., R. C. Kingsbury, and S. L. McKnight. 1988. 'Functional dissection of VP16, the trans-activator of herpes simplex virus immediate early gene expression', Genes Dev, 2: 718-29.
Wang, Y., J. Xu, T. Pierson, B. W. Omalley, and S. Y. Tsai. 1997. 'Positive and negative regulation of gene expression in eukaryotic cells with an inducible transcriptional regulator', Gene Therapy, 4: 432-41.
Witzgall, R., E. O'Leary, A. Leaf, D. Onaldi, and J. V. Bonventre. 1994. 'The Kruppel-associated box-A (KRAB-A) domain of zinc finger proteins mediates transcriptional repression', Proceedings of the National Academy of Sciences of the United States of America, 91: 4514-8.
Wu, C. Y., K. T. Roybal, E. M. Puchner, J. Onuffer, and W. A. Lim. 2015. 'Remote control of therapeutic T cells through a small molecule-gated chimeric receptor', Science, 350: aab4077.
