
Gadusol is a multi-hydroxyl ketone, which is converted from 7-phosphosedoheptulose (S7P) by isomerization, methylation and dehydrogenation. Generally, for simple enzyme-catalyzed reactions, we can choose a suitable kinetic equation for the enzymatic reaction to simulate. However, sedoheptulose, the precursor substances of gadusol, is not a useable fermentation raw materials, due to its high price, low yield and lack of direct kinase. Therefore, considering S7P is an intermediate product of metabolism, we attempted to produce gadusol directly by using intracellular S7P as a substrate.
Most enzymes in yeast cells are tightly regulated, and their kinetic equations are complex and lack of experimental data. In the same time, we are not sure of the kinetic mechanism of EEVS and MT-Ox. Therefore, we decided to use an indirect approach to analyze the yield of gadusol.
All in all, we chose Metabolic Flux Analysis (MFA) model to evaluate precursor supplement and to guide our wet lab optimization.

Fig.1 The Main Parts Of The HMP & EMP Metabolic Pathways In Yeast Cells, From KEGG
We simplified some continuous reactions without branches into one step, and we only took the carbon number of different saccharides into account. because saccharides with same carbon number here can transform into one another reversibly. (C6: Glucose-6-phosphate & Fructose-6-phosphate, C5: Xylulose-5-phosphate & Ribose-5-phosphate & Ribose-5-phosphate, C4: Erythrose-4-phosphate, C3: Glyceraldehyde-3-phosphate & Dihydroxyacetone phosphate)

Fig.2 Simplified Metabolic Pathways(Tkl2 Is Skipped, The Same Below)
Obviously, the generation rate of gadusol equals to the rate difference of Tkl1 and Tal1. To determine the values of Tkl1 and Tal1's flux, we chose the following metabolic rates with clear biological meanings:
u : nucleotide synthesis
p : aromatic amino acid synthesis
r : sum of all metabolic fluxes of endogenous 3-Phospho-glyceraldehyde
s : rate of polysaccharide synthesis
o : rate of intracellular NDAPH consumption
i : rate of glucose intake
They worked as the basic rates. (here we use relative flux so their unit is not shown, however, have the dimension of mol/(L*H))
After discussion with the wet lab group, we made a hypothesis based on theory of Microbial Transformation by Resting Cell:
1. The metabolic rate of yeast cells does not change significantly after the introduction of exogenous genes.
2. No accumulation of metabolic intermediates occurs after the introduction of the exogenous gene.
3. We only consider NADPH as the sole reducing power, skips NADH and ATP (energy related)

Fig.3 Simplified Metabolic Pathways & Related Calculation
After arithmetic operations, we obtained the relation between gadusol synthetic rate and the basic rates (include u, p, r, s, i, o).
12i-o-6r-10u-12s-8p=18g
However, it's easy to point out that there will be 12i-o-6r-10u-12s-8p=0 for chassis cells considering the absence of S7P. Therefore, if the basic rates remain constant, gadusol will not accumulate. We proposed that the accumulation of gadusol mainly depends on the rate of glucose absorption rate i, and other basic rates remained constant. otherwise it might cause metabolic imbalances.
Introducing a new carbon sourece to enhance the flux of the HMP pathway is the most direct way. After comparison, we finally chose a cheap carbon source xylose, which can not only effectively improve sedoheptulose-7-Phosphate supplement, but also have a good economic potential.
Xylose is isomerized to xylulose by reduction and oxidation, and subsequently converted to X5P by xylulose kinase. The xylose pathway consumes NADPH, the import of this pathway would additionally enhance the pathway GDH (C6-C5).

Fig.4 Simplified Metabolic Pathways After Introducing Xylose Utilization Pathway
After introducing that,there is an equation: 12i+9x-o-6r-10u-12s-8p=18g
Also, we found that xylose can replace the role of glucose to support the cell living, though its efficiency might be lower than glucose, and the low efficiency of Tal1 may be a difficulty to meeting the needs, so we believe that a dual-substrate mixed fermentation and subsequent optimizations are needed.
Cutting of the other consumption of S7P could be another way to increasing the production of gadusol. Besides, previous studies (Transaldolase deficiency: report of 12 new cases and further delineation of the phenotype) hinted that knocking out of Tal1 will not cause yeast cell death. However, while the accumulation of S7P, it might make the HMP pathway less efficient.
If we knock out Tal1, the accumulated S7P will be forced to flow to gadusol. There are 3g=o-2u+2p and 12i=-2u+12s+20p+6r+7o. We found that gadusol yield and glucose intake could theoretically be determined if basic rates were known.

Fig.5 Simplified Metabolic Pathways After Knocking Out Transaladolase Gene
Combining the two methods, we have 3g=o-2u+3x+2p, 12i=9x-2u+12s+20p+6r+7o. Unlike the introduction of xylose pathway alone, here xylose intake must be accompanied by glucose intake, thus the i value is always greater than 0. At the same time, the value of i and x must satisfy the above equation to get the maximum yield, otherwise it will cause the accumulation of intermediates or incomplete absorption of carbon sources.

Fig.6 Simplified Metabolic Pathways After Knocking Out Transaladolase Gene & Introducing Xylose Utilization Pathway
Based on article Kinetic Modeling of Saccharomyces cerevisiae Central Metabolic Network, we assumed that u=3, p=3, o=20, s=10, r=150, and then calculated their effect.

Fig.7 a) Yeast Without Optimization; b) Introducing Xylose Utilization Pathway; c) Introducing Xylose Utilization Pathway & Knocking Out Tal1 (While Xylose intake=0, it is the point only knocking out Tal1)
1. Xylose is less efficiency than glucose (1 g xylose equal to 0.9 g glucose), however, it can strengthen HMP'flux , is less regulated and can be absorbed along with glucose so it can be added as an extra carbon source. While glucose intaking has met the cells' living, xylose intaking will be transformed into gadusol in a high ratio.
2. Knocking out Transaldolase may couple cell growth with gradual generation, it will always lead to a yield over 0 ( point xylose intake=0 in fig.7c ), although is not too high.
3. Combining two ways can enhance HMP while cut off carbon flux S7P → E4P. Glucose can meet the cells' need, and xylose can be transformed into gadusol. It has a higher efficiency than only introduce a xylose pathway. Besides, by doing this, glucose and xylose intaking will be related, thus, there is a best glucose-xylose ratio to make the gadusol yield highest.
These three optimization plan works in different way, but they can both enhance yield of gadusol. So we decided to introduce a xylose utilization pathway first to ensure the HMP pathway reliable enough to support subsequent molecular operations. Then, we plan to further knock out Tal1 of the xylose utilization strain. We will detect the yield obtained by different ways, compared to the simulations in this section, perform model optimization, and finally simulate a more precise yield range.
When we introduced the MT-Ox/EEVS expression vector and xylose utilization pathway genes vector into the cell after they were constructed, we found something unexpected: the growth and metabolic rates of the yeast changed apparently, the OD600 value reach around 15 in YNB-based glucose media (see Experiments section). Therefore, it is necessary for us to make a more reliable model.
Firstly, we found a simpler way of calculation. For a metabolic network with several reactions (n) involving several substances(m) (Ps. Substances can enter or exit cells freely are not considered), it can be written as an m*n matrix, in which each column is a reaction, each row is a substance, and the matrix element [i, j] is the stoichiometric number of the substance i in reaction j. The principle of "production is positive, consumption is negative" is followed. Also we set a column vector v, vector vi is the reaction rate of the ith reaction. For any substance i, the sum of the rates of all relevant reactions times its reaction stoichiometry number should be equal to 0 according to the assumption of non-accumulation.
According to the above description,it's easy to show in a mathematical way: for any substance i ,we have equation 1, and for a matrix Mm*n consisting of m substances, there should be equation 2.
 Equation 1
Equation 1
 Equation 2
Equation 2

Fig.8 An Example Of The Method
Secondly, we carefully investigated the metabolic pathways of yeast by books and databases such as KEGG and NCBI, compiled a more detailed reaction table and optimized some unnecessary reactions for our reaction conditions, which are also labeled with imaginary line in the figure.
Compared with the previous method, the metabolic pathway here is relatively more complete, and the consideration of NAD and ATP has been added, we also considered the growth of cells. (here we skipped some reactions and metabolisms that is not important, and simplified non-branch pathways)

Fig.9 MFA Of Yeast Grew On Glucose Medium
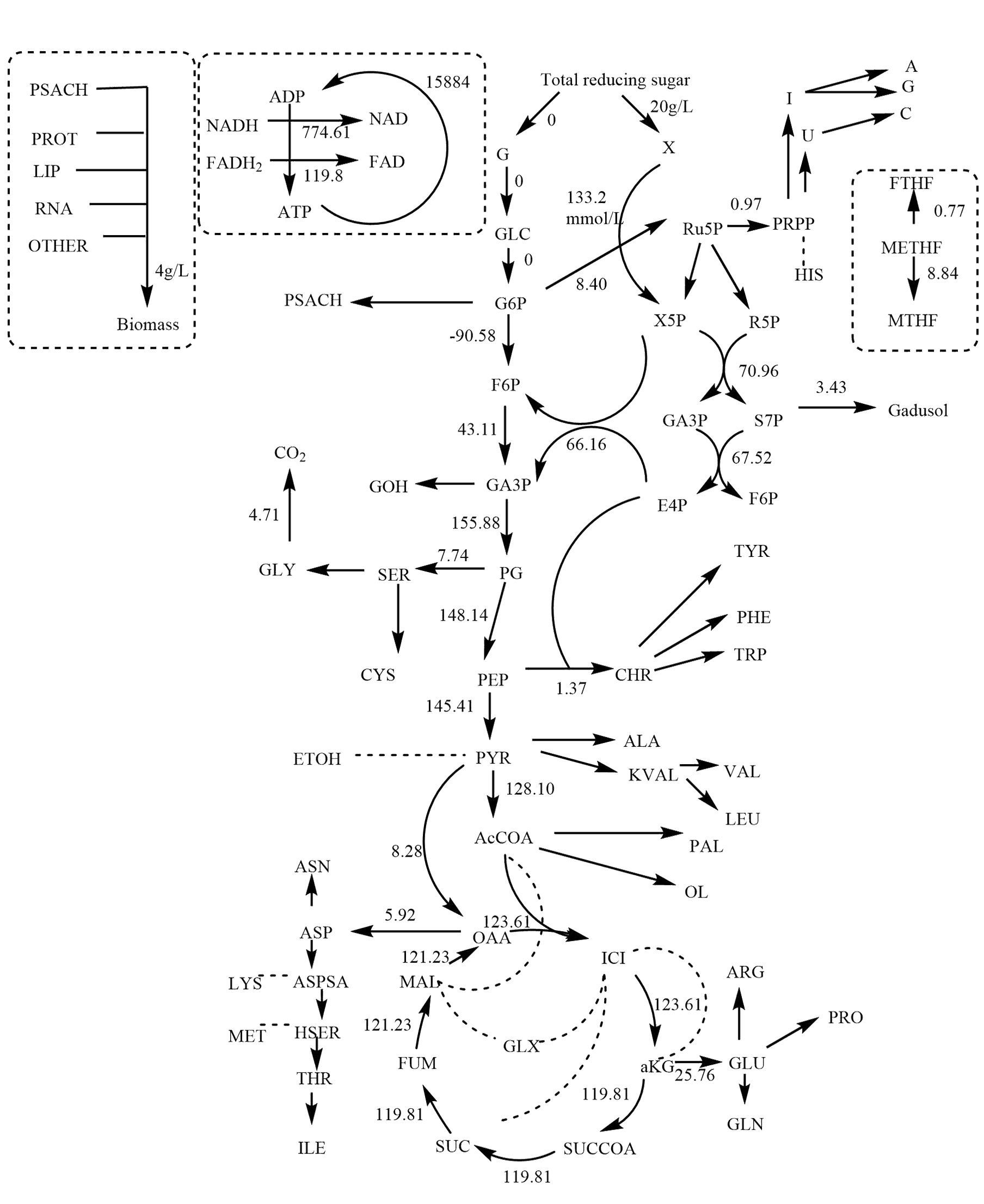
Fig.10 MFA Of Yeast Grew On Xylose Medium
From the wet lab data, we assumed that: The lowest ATP no-load consumption rate: 1500; Minimum cell generation: 2 g/L.
To estimate the effect of the three previously mentioned methods on the gadusol yield, we only performed maximizing by linear programming to determine the range of values for the product acquisition. (The way of solving is in the appendix.)
For us, we hope the cells to produce as much gadusol as possible, however, yeast obviously wants to generate more progeny on its own.
Therefore we use two approaches to estimate the gadusol yield:
1: Find the maximum amount of gadusol generated as an upper limit;
2: Find the yield of gadusol for the maximum growth of the yeast as a lower limit.
Finally, our predicted gadusol yield is between upper limit and lower limit in Fig.11 below.

Fig.11 Yield Range Prediction & Experimental Data ED=Experimental Data, PDR=Predicted Data Range
Optimization 1: Introducing Xylose Utilization Pathway; Optimization 2: Knocking Out Transaldolase; Optimization 3: Combine Them Together
At the same time, we pointed out:
1. The intake of xylose gave a lower yield than the intake of glucose, but their yield range was both broad, it could not give an accurate prediction result.
2. Knockdown of Tal1 might cause cell growth coupled to gadusol production, which lower the upper limit but greatly increase the lower limit. Besides, it also made glucose and xylose intake correlate. In opt.3, the best glucose-xylose ratio is 17.7:2.3 .
3. Here we didn’t choose the minimum gadusol yield in linear programming as a lower limit, the reason is that, if we choose minimum gadusol, it will cause an abnormally low efficiency of carbon source using(e.g. all carbon is used to burn and all ATP consumed by no-load cycling), and get a meaningless result. This is the limit of linear programming and this model.
According to the prediction: knocking out Tal1 may not cause the death and may enhance the flux towards gadusol. Since we have introduced the xylose utilization pathway into the yeast and gain a great ability of cell growth & gadusol production, we plan to knock out the Tal1 to further increase its production. According to the data of simulation,it may increase 2 times at least.
For this model, what we want to do next is to find out the exact flux maximum of key reactions and to reduce predication errors by further integrated wet lab results and to stimulate other molecular engineering operations on production and cell growth, such as overexpressing Transketolase (Tkl1), and further guiding experiments.




