Introduction to our Model
One of the most challenging aspects of metabolic engineering is the size and complexity of any given biological system. Metabolic modelling in silico is one of the solutions to this challenge, as it provides insight into these biological systems, even on large genome-scale models [1]. The premise of our work is the biodegradation of microplastics, specifically polyethylene terephthalate (PET) and polyethylene (PE) by microorganisms such as Bacillus subtilis. This organism has been shown to have the capacity of degrading these microplastics in small amounts [2]. Our goal is to use metabolic engineering to increase the biodegradation capabilities of B. subtilis.
In order to achieve this, a biological model was used to predict the best targets for metabolic engineering, via knockout (KO) or under/over expression (UO). The model iBsu1103 [3] was chosen, a stoichiometric model with 1103 genes. This model did not include the degradation pathways for PET and PE, therefore it was necessary to insert the reactions constituting these pathways into the model. We resorted to Optflux [4], a metabolic engineering software, for the simulation and optimization (target prediction) of the model. Our model can be downloaded here.
To summarize, our goals for the modelling and informatics team were the following:
1) Insert PET and PE pathways into the iBsu1103 model;
2) Check for cell growth in environments with PET or PE as the sole carbon source;
3) Optimize innocuous metabolite (ethanol) and biomass production with PET or PE as the sole carbon source (find targets for metabolic engineering).
Degradation of PET
Polyethylene terephthalate (PET) is a chemical polymer formed by purely synthetic processes. One of those involves ethylene glycol (EG) and terephthalic acid (TPA) forming bis(hydroxyethyl) terephthalate (BHET) which is then polymerised into the final product [5]. It is highly used due to its durability, resistance and non-degradable nature [6]. From all the solid waste produced, PET is responsible for 12 % of its volume [7], and the most abundant polyester plastic [8], being a current threat to the marine environment and human health.
There are chemical recycling solutions, but their high costs and secondary products make them less appealing [9,10], so a new way to remove these microplastics from the environment is needed. In 2016, Yoshida et al. [11] described a bacterium - Ideonella sakaiensis - capable of degrading PET and using it as its main carbon source. This bacterium secretes two enzymes to the extracellular medium: poly(ethylene terephthalate) hydrolase (PETase) that converts PET into mono(2-hydroxyethyl) terephthalic acid (MHET) and MHETase that further hydrolyses MHET into TPA and EG [11] (Figure 1). Afterwards, it absorbs TPA and EG, utilizing them as carbon sources.

Figure 1: Schematic representation of Polyethylene Terephthalate (PET) degradation pathway performed by Ideonella sakaiensis. PETase, polyethylene terephthalate (PET) hydrolase; BHET, bis(2-hydroxyethyl) terephthalic acid; MHET, mono(2-hydroxyethyl) terephthalic acid; TPA, terephthalic acid; EG, Ethylene glycol [12].
Bacillus subtilis has shown the capacity to degrade PET [2,12], but the exact mechanism is unknown. Since the pathway from Ideonella sakaiensis has shown to be the most effective one to degrade PET, we decided to express the two enzymes in B. subtilis, therefore we inserted this pathway on our B. subtilis model (iBsu1103) as well. Then, we needed to simulate how using PET as a carbon source would impact Bacillus subtilis metabolism. Afterwards, several optimizations were executed in order to find target reactions for metabolic engineering, with the objective of maximizing biomass and the production of a naturally occurring metabolite (ethanol).
Degradation of PE
Several studies have reported the potential polyethylene (PE) degradation capabilities of Bacillus subtilis [13]. This organism was found to be able to degrade a wide range of PE variants when applied as an isolated culture or as part of a broader microbial consortium [14-17]. Although the PE degradation capability of B. subtilis has been fairly well reported, few studies objectively define the underlying biochemical pathways and genes involved in the process. It is thought that this gram-positive bacterium employs a similar degradation strategy to that of other well-reported organisms such as Rhodococcus sp. and Pseudomonas sp. [18].

Figure 2: Aerobic (in green background) and anaerobic (in red background) alkane degradation pathway [17].
Bacillus subtilis degrades n-alkanes by adding a terminal carboxyl group through a series of oxidation steps. Once carboxylated, n-alkanes are effectively converted into fatty acids and these metabolites follow the β-oxidation pathway to be completely oxidized into CO2. An alkane hydroxylase - alkane monooxygenase - is thought to be the key initiator in the terminal oxidation of n-alkanes into primary alcohols; these are then further oxidized into aldehydes and carboxylic acids, by the alcohol dehydrogenase and aldehyde dehydrogenase, respectively. Fatty acids are incorporated into the citric acid cycle and through the long-chain-fatty-acid-CoA ligase, they generate ATP and reductive power [19-22].
This knowledge was the underlying premise in our project to construct a Bacillus subtilis chassis (Genetically Modified Organism, GMO) metabolic model capable of degrading PE.
Environmental Conditions and WildType Simulation
Our aim is to develop a bacteria that can feed off microplastics, so we started by evaluating the possibility of the cell using PET as the only carbon source by simulating the cell growth in these conditions. To do so, we restricted the entrance of several metabolites into the cell, including glucose, only allowing the input of certain indispensable metabolites and PET or PE, depending on the carbon source being tested. These environmental conditions can be seen in Figure 3, where the “ins and outs” of metabolites are limited or completely blocked. PET and PE intake is limited to better simulate environmental conditions.

Figure 3: Environmental conditions.
We performed a WT simulation using environmental conditions where PET was the only carbon source, maximizing biomass as the objective function using the parsimonious Flux Balance Analysis (pFBA) as the simulation method, as shown in Figure 6. This method seeks to “minimize the flux associated with each reaction in the model while maintaining optimum flux through the objective function” [23]. The biomass value was 0.9048 (Figure 4), indicating growth while using only PET as carbon source.

Figure 4: Simulation results for the medium that only contains PET as the carbon source.
Afterwards, we performed a WT simulation to check if there was cell growth while using only PE as the carbon source, using the same method as with PET. Unfortunately, the simulation shows no biomass while feeding solely on PE, indicating that a detailed analysis of the pathways and all its precursors needs to be done.

Figure 5: WildType simulation using only PE as carbon source.

Figure 6: WildType simulation menu using only PET as carbon source.
Biomass and Ethanol Optimization Results
With the simulations indicating cell growth in environments with only PET as carbon source, we now intend to maximize PET degradation flux by converting it into another metabolite that could have a commercial value, while keeping the cells viable. We decided to maximize ethanol production by an evolutionary algorithm known as SPEA2 (Strength Pareto Evolutionary Algorithm 2), a method consisting of parent generations of candidate solutions, leaving their best “traits” for future generations to inherit, constantly increasing objective function values.
The objective function used for this optimization was biomass-product coupled yield (BPCY) with the product being the reaction that represents ethanol drain from the system:

Optflux allows the user to do this optimization for both KO and UO sets of solutions. In this case we performed the optimizations using the linear minimization of metabolic adjustment (LMOMA) approach as shown in Figure 7. LMOMA is a simulation method that minimizes repercussions caused by KO or UO in the redistribution of metabolic fluxes regarding the ones from the wild type.

Figure 7: Perform Strain Optimization using the only PET medium and maximizing biomass and ethanol production.
After several rounds of optimizations, we obtained various solutions for maximizing ethanol production while consuming PET as carbon source. An example solution is shown in Figure 8. However, since using PE as a single carbon source did not support the cell growth, it was not possible to proceed to the optimization process.
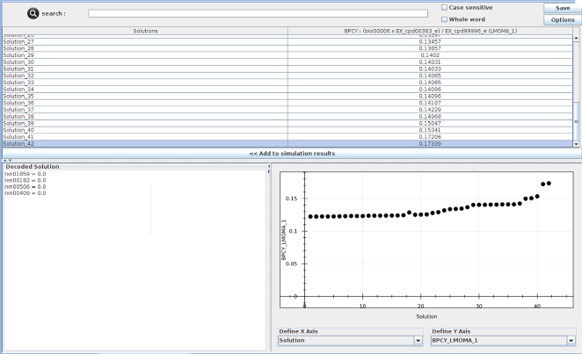
Figure 8: Example of best optimization solutions gotten from Optflux.
Our best result for Knockout optimizations had a biomass growth rate of 0,6619 h-1 and a flux of ethanol of 2,6195 mmol.(gDW.h)-1. For the UO optimization, the optimum solution led to a slower growth rate when compared with the KO solution (0.4066 h-1), but the flux of ethanol was more than 2-fold higher 6.4448 mmol.(gDW.h)-1. The most promising optimization results are shown in Table 1.
Table 1: Simulation and optimization results.

The optimization results from ID2 (see Table 1) identified 2 possible reactions suited for gene knockout, those of 1,6-Diphosphofructose aldolase (E.C. 1.2.1.10, rxn00786) and acetaldehyde dehydrogenase (E.C. 4.1.2.13, rxn00506). As it is already known PET degradation results in the production of two major metabolites, TPA and monoethylene glycol, which are incorporated into the cell. Monoethylene glycol is readily converted into oxaloacetate, resulting in an overburden of this metabolite which in normal circumstances is converted into energy and reductive power in the Krebs cycle. Instead, excess oxaloacetate is redirected to the second stage of glycolysis (conversion into phosphoenolpyruvate) which can, in turn, explain the overproduction of ethanol. The knockout of 1,6-Diphosphofructose aldolase would inhibit an important step in the catabolization of sugars, which is justified since they are the major sources of carbon and end up being the catabolites of PET. Moreover, acetaldehyde dehydrogenase knockout would limit, to some extent, the reentrance of oxaloacetate catabolites in the Krebs cycle effectively redirecting them to the production of ethanol.
The optimization results from ID3 indicate 3 candidate reactions for gene knockout; those of acetaldehyde dehydrogenase (E.C. 4.1.2.13, rxn00506), threonine synthase (E.C. 4.2.3.1, rxn 01069), and glycolaldehyde transferase (E.C. 2.2.1.1, rxn00785). The reasoning behind acetaldehyde dehydrogenase knockout has been presented before for the ID2 solution. The knockout of glycolaldehyde transferase would lead to a decrease in the flux between glycolysis and the pentose phosphate pathway, which is manifested by a decrease in D-glyceraldehyde 3-phosphate flux. This process results in an inhibition of the second stage of glycolysis when PET is the sole source of carbon. The knockout of threonine synthase will effectively isolate the glyoxylate metabolism from other important metabolisms (e.g. aspartate, alanine, glutamine, lysine, and arginine) which could, in turn, redirect oxaloacetate to the production of ethanol.
The optimization results from ID4 indicate 3 possible reactions for gene knockout; those of adenine nucleoside phosphorylase (E.C. 2.7.4.6, rxn00409), glutamate dehydrogenase (E.C. 1.4.1.2, rxn00182), and nucleoside-diphosphate kinase (E.C. 2.4.2.1, rxn01859). The knockout of adenine nucleoside phosphorylase and nucleoside-diphosphate kinase isolates the glyoxylate metabolism from other important purine, pyrimidine, and aminoacid metabolisms (e.g. aspartate, alanine, glutamine, etc.) which could, in turn, redirect oxaloacetate to the production of ethanol. Aldehyde dehydrogenase (E.C:1.2.1.3, rxn00506,) knockout would inhibit acetate production, which could lead to higher growth rates. Moreover, the knockout of glutamate dehydrogenase would inhibit the formation of glutamate and conserve oxaloacetate for ethanol production.
In the solution ID5, the strong upregulation of CTP synthase (E.C. 6.3.4.2, rxn 00410, 32.0) and downregulation of nucleoside-diphosphate kinase (E.C. 2.4.2.1, rxn01859, 0.03125) favor the formation of methymalonate and R-3-amino-isobutanoate into pyridine. The extra flux of these metabolites is then converted into acetyl-CoA (valine, leucine, and isoleucine metabolism) and redirected to ethanol production. Furthermore, the downregulation of 3-hydroxyacyl-CoA dehydrogenase (E.C. 1.1.1.35, rxn 03249, 0.125), buffer the glyoxylate metabolism from the purine, pyrimidine, and amino acid metabolisms (e.g. aspartate, alanine, glutamine, etc.) which can, in turn, redirect oxaloacetate to the production of ethanol.
In the solution ID6, the upregulation of aspartate kinase (E.C. 2.7.2.4, rxn00337, 8.0) and downregulation of serine-pyruvate transaminase (E.C. 2.6.1.51, rxn00422, 0.03125) buffer the glyoxylate metabolism from other amino acid metabolisms (e.g. aspartate, alanine, glutamine, lysine, proline, and arginine), which could in turn redirect oxaloacetate to the production of ethanol.
These findings indicate possible gene targets either for KO or UO that are suited for increasing ethanol production from PET. Nevertheless, these results should be further analyzed before in vivo implementation since some potential gene candidates are critical to the vital cell processes and metabolic engineering approaches should be applied with caution (e.g adenine nucleoside phosphorylase interferes with RNA and DNA synthesis) in order to not compromise the biomass growth.
Conclusion
Our results show that Bacillus subtilis can use PET as their sole carbon source and use it to produce native metabolites such as ethanol. Unfortunately, the PE simulations did not present good results and so it was impossible for us to computationally analyze its degradation benefits to the cell. One possible explanation for these results is that the cell might not have been able to link the pathway to its native metabolism thus resulting in unfavourable accumulation of unwanted metabolites. Nevertheless, we envision that these in silico results could support the wet lab experiences by providing possible gene targets for further metabolic engineering strategies, either gene knockouts or under/over expressions. We hope that these findings could save time in the lab by narrowing the potential gene targets for metabolic engineering and by understanding the respective impact on the cell’s metabolism.
The final goal of our project is to develop a B. subtilis strain capable of efficiently degrading PE - by its native pathways - and PET - using heterologous I. sakaiensis enzymes - into innocuous metabolites. The results obtained support the possibility of developing a Bacillus strain able to, at least, grow using PET as the sole carbon source, converting it to ethanol. In a more advanced state of our project, we hope to link the PET and PE degrading metabolism with the formation of a high added value compound (e.g. surfactin). This engineered strain would increase the potential of applying our solution in real life since it helps to diminish the microplastics problem and, in parallel, produces a high added value compound in a sustainable fashion.
References
- C. Gu, G.B. Kim, W.J. Kim, H.U. Kim, S.Y. Lee, Current status and applications of genome-scale metabolic models, Genome Biol. 20 (2019) 121. https://doi.org/10.1186/s13059-019-1730-3.
- J. Oliveira, A. Belchior, V.D. da Silva, A. Rotter, Ž. Petrovski, P.L. Almeida, N.D. Lourenço, S.P. Gaudêncio, Marine Environmental Plastic Pollution: Mitigation by Microorganism Degradation and Recycling Valorization, Front. Mar. Sci. 7 (2020). https://doi.org/10.3389/fmars.2020.567126.
- C.S. Henry, J.F. Zinner, M.P. Cohoon, R.L. Stevens, iBsu1103: A new genome-scale metabolic model of Bacillus subtilis based on SEED annotations, Genome Biol. 10 (2009) 1–15. https://doi.org/10.1186/gb-2009-10-6-r69.
- I. Rocha, P. Maia, P. Evangelista, P. Vilaça, S. Soares, J.P. Pinto, J. Nielsen, K.R. Patil, E.C. Ferreira, M. Rocha, OptFlux: an open-source software platform for, BMC Syst. Biol. 4 (2010). https://doi.org/https://doi.org/10.1186/1752-0509-4-45.
- F. Awaja, D. Pavel, Recycling of PET, Eur. Polym. J. 41 (2005) 1453–1477. https://doi.org/10.1016/j.eurpolymj.2005.02.005.
- Z. Leng, R.K. Padhan, A. Sreeram, Production of a sustainable paving material through chemical recycling of waste PET into crumb rubber modified asphalt, J. Clean. Prod. 180 (2018) 682–688. https://doi.org/10.1016/j.jclepro.2018.01.171.
- A.M. Atta, A.F. El-Kafrawy, M.H. Aly, A.A.A. Abdel-Azim, New epoxy resins based on recycled poly(ethylene terephthalate) as organic coatings, Prog. Org. Coatings. 58 (2007) 13–22. https://doi.org/10.1016/j.porgcoat.2006.11.001.
- V. Tournier, C.M. Topham, A. Gilles, B. David, C. Folgoas, E. Moya-Leclair, E. Kamionka, M.L. Desrousseaux, H. Texier, S. Gavalda, M. Cot, E. Guémard, M. Dalibey, J. Nomme, G. Cioci, S. Barbe, M. Chateau, I. André, S. Duquesne, A. Marty, An engineered PET depolymerase to break down and recycle plastic bottles, Nature. 580 (2020) 216–219. https://doi.org/10.1038/s41586-020-2149-4.
- M.H. Vakili, M.H. Fard, Chemical Recycling of Poly Ethylene Terephthalate Wastes, Ind. Eng. Chem. Res. 8 (2010) 839–846. https://doi.org/10.1016/j.polymdegradstab.2009.11.026
- S. Matsumura, A.R. Hlil, C. Lepiller, J. Gaudet, D. Guay, Z. Shi, S. Holdcroft, A.S. Hay, Stability and Utility of Pyridyl Disulfide Functionality in RAFT and Conventional Radical Polymerizations, J. Polym. Sci. Part A Polym. Chem. 46 (2008) 7207–7224. https://doi.org/10.1002/pola.
- Y. Yang, J. Yang, L. Jiang, Comment on "a bacterium that degrades and assimilates poly(ethylene terephthalate) ", Science (80-. ). 353 (2016) 759. https://doi.org/10.1126/science.aaf8305.
- N. Mohanan, Z. Montazer, P.K. Sharma, D.B. Levin, Microbial and Enzymatic Degradation of Synthetic Plastics, Front. Microbiol. 11 (2020). https://doi.org/10.3389/fmicb.2020.580709.
- P.P. Vimala, L. Mathew, Biodegradation of Polyethylene Using Bacillus Subtilis, Procedia Technol. 24 (2016) 232–239. https://doi.org/10.1016/J.PROTCY.2016.05.031.
- B. Nowak, J. Pajak, M. Drozd-Bratkowicz, G. Rymarz, Microorganisms participating in the biodegradation of modified polyethylene films in different soils under laboratory conditions, Int. Biodeterior. Biodegradation. 65 (2011) 757–767. https://doi.org/10.1016/J.IBIOD.2011.04.007.
- C. Abrusci, J.L. Pablos, T. Corrales, J. López-Marín, I. Marín, F. Catalina, Biodegradation of photo-degraded mulching films based on polyethylenes and stearates of calcium and iron as pro-oxidant additives, Int. Biodeterior. Biodegradation. 65 (2011) 451–459. https://doi.org/10.1016/J.IBIOD.2010.10.012.
- K. Harshvardhan, B. Jha, Biodegradation of low-density polyethylene by marine bacteria from pelagic waters, Arabian Sea, India, Mar. Pollut. Bull. 77 (2013) 100–106. https://doi.org/10.1016/J.MARPOLBUL.2013.10.025.
- Z. Montazer, M.B.H. Najafi, D.B. Levin, Microbial degradation of low-density polyethylene and synthesis of polyhydroxyalkanoate polymers, Https://Doi.Org/10.1139/Cjm-2018-0335. 65 (2018) 224–234. https://doi.org/10.1139/CJM-2018-0335.
- C. Park, W. Park, Survival and energy producing strategies of Alkane degraders under extreme conditions and their biotechnological potential, Front. Microbiol. 9 (2018) 1–15. https://doi.org/10.3389/fmicb.2018.01081.
- Z. Montazer, M.B.H. Najafi, D.B. Levin, Challenges with verifying microbial degradation of polyethylene, Polymers (Basel). 12 (2020). https://doi.org/10.3390/polym12010123.
- S. Ghatge, Y. Yang, J.H. Ahn, H.G. Hur, Biodegradation of polyethylene: a brief review, Appl. Biol. Chem. 63 (2020). https://doi.org/10.1186/s13765-020-00511-3.
- S.E. Diomandé, C. Nguyen-The, M.H. Guinebretière, V. Broussolle, J. Brillard, Role of fatty acids in Bacillus environmental adaptation, Front. Microbiol. 6 (2015) 1–20. https://doi.org/10.3389/fmicb.2015.00813.
- M.L. Jenior, T.J. Moutinho, B. V. Dougherty, J.A. Papin, Transcriptome-guided parsimonious flux analysis improves predictions with metabolic networks in complex environments, PLoS Comput. Biol. 16 (2020). https://doi.org/10.1371/journal.pcbi.1007099.
- D. Segrè, D. Vitkup, G.M. Church, Analysis of optimality in natural and perturbed metabolic networks, Proc. Natl. Acad. Sci. U. S. A. 99 (2002) 15112–15117. https://doi.org/10.1073/pnas.232349399.







