Results
Goal:
- Construct dual plasmid system in bacteria capable of producing α-terpineol
Approach:
After doing literature research, we discovered a series of pathways that if connected through enzymatic activity had the capability of producing α-terpineol. The first pathway is known as the yeast Mevalonate (MEV) pathway. This pathway starts with acetyl-CoA as its substrate and though a series of enzymatic conversions produces IPP & DMAPP. The second pathway is the basic monoterpene production pathway. This pathway is able to take IPP & DMAPP and convert these into α-terpineol via a two-step conversion utilizing geranyl pyrophosphate synthase (GPPS) & α-terpineol synthase (αTS) as shown in the figure below. We used the αTS from the organism Vitis vinifera in the following experiments, hereafter referred to as VvTs.
*The αTS used in the following experiments is referenced as VvTs as this αTS came from the organism Vitis vinifera*

In our dual plasmid model, the first plasmid was readily available via Addgene known as JBEI-3085 (#87950). This plasmid contains the enzymes found in the yeast MEV pathway. The second plasmid also known as pTrc_GppsGB_VvTs had to be constructed to contain a GPPS & the Vitis vinifera αTS (VvTs). In order to accomplish this, we were able to digest and isolate a backbone from our source plasmid (pTrc_GPPS_LS) via gel purification that has been previously used for other monoterpene production as shown in the figure below. The 4.1kb backbone (indicated in the red box) was successfully gel-purified from the excised 2.6kb insert (bottom band).

Gel purification of backbone
After successful gel purification of our desired backbone, we performed NEBuilder HiFi DNA assembly with our designed insert and sequentially transformed this plasmid (pTrc_GppsGb_VvTs) into competent NEB 5-α cells.

Transformation of pTrc_GppsGb_VvTs into NEB 5- α cells
A miniprep kit was used to extract and purify our constructed plasmid for transformation into MG1655 E. coli cells. To confirm cloning of our plasmid we conducted a restriction enzyme digest and sent off our plasmid for sequencing. Once confirmed, we double transformed pTrc_GppsGb_VvTs along with JBEI-3085. A restriction enzyme digest was also used to confirm these transformations as shown in the figure below.
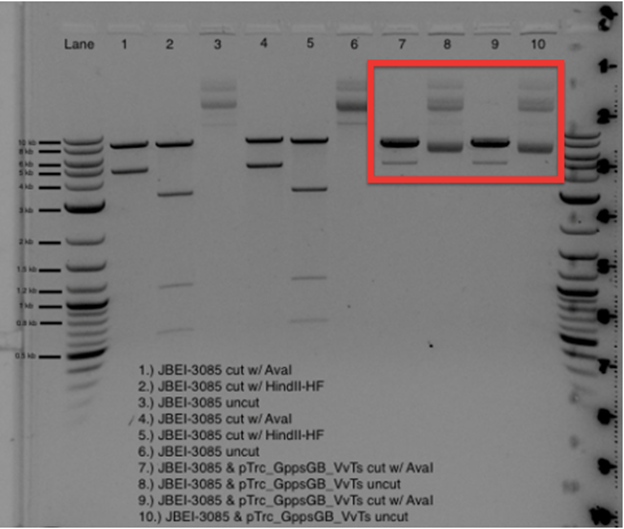
Verification of double transformation of pTrc_GppsGb_VvTs & JBEI-3085 into MG1655 cells
With the confirmation of these transformations, we were able to proceed with production trials of α-terpineol. The production assays were ran by inducing our bacterial cultures with IPTG in glucose rich media known as EZ-rich media with a dodecane overlay followed by an incubation period of 72 hours while shaking at 250rpm. After induction, any α-terpineol produced should be in the organic dodecane layer. The samples were extracted by adding ethyl acetate and removing only the top organic layer for GC/MS quantification.
Before production trials were run, the extraction method and limit of detection for our GC/MS approach were evaluated. The extraction method was verified by spiking in a known mount of α-terpineol to blank samples. The samples were then extracted and sent for GC/MS analysis as shown by the image below.

The image above is a control sample containing α-terpineol spiked in (before extraction with ethyl acetate). A clear peak for α-terpineol can be seen at the timepoint of 18.40. The level of detection was determined by spiking in known amounts of α-terpineol in increasing concentrations and creating a calibration curve as shown below. Using this method, we were able to detect a spiked in concentration down to 10 - 15mM α-terpineol.

Following the confirmation of the extraction method and level of detection, we were able to begin our production runs of α-terpineol. Sample data is shown below.

Above is an image from a production run of one of our samples which indicated no detection of α-terpineol. Based on the data above, it is inconclusive as to whether our system is producing α-terpineol. It is possible our system is producing α-terpineol at a level below the detection limit by GC/MS. Further tests and optimizations need to be conducted to clarify the results.

