
Cycle 1 - Biosynthesis of Variecolin
Design – Search for the biosynthetic genes of variecolin
The first step of this project comprised the search for relevant gene clusters responsible for variecolin production, utilizing bioinformatic tools. In silico analyses revealed that Aspergillus aculeatus ATCC 16872 (CBS 172.66) contains a putative biosynthetic gene cluster for variecolin, which encodes the chimeric terpene synthase consisting of prenyltransferase (PT) and terpene cyclase (TC) domains, the cytochrome P450 monooxygenase, and the ATP-binding cassette (ABC) transporter. The biosynthetic pathway of variecolin can be predicted as follows: The PT domain of the terpene synthase first catalyzes the condensation of one molecule of dimethylallyl pyrophosphate (DMAPP) and four molecules of isopentenyl pyrophosphate (IPP), to form a C25 geranylfarnesyl pyrophosphate (GFPP). The TC domain then cyclizes GFPP, to afford a hydrocarbon intermediate with a 5-8-6-5-fused ring system, which undergoes multiple cycles of oxidative reactions catalyzed by the P450 to produce variecolin.

Figure 1. Predicted biosynthetic pathways of variecolin.
For heterologous production of variecolin, Aspergillus oryzae NSAR1 (niaD-, sC, ΔargB, adeA)[1] was used as an auxotrophic fungal host. A terpene synthase (TS) gene was introduced into the pAdeA vector that contains the adeA prototrophic marker for adenine synthesis, while diverse P450 gene constructs were prepared using the pTAex3 vector that harbors the argB prototrophic gene marker. The plasmids, pAdeA-TS and pTAex3-P450, were transformed into the auxotrophic A. oryzae NSAR1 recipient strain for variecolin production.
Test – Analysis of the product - variecolin
HPLC and NMR analyses were performed to verify whether our biosynthetic approach of variecolin was successful.
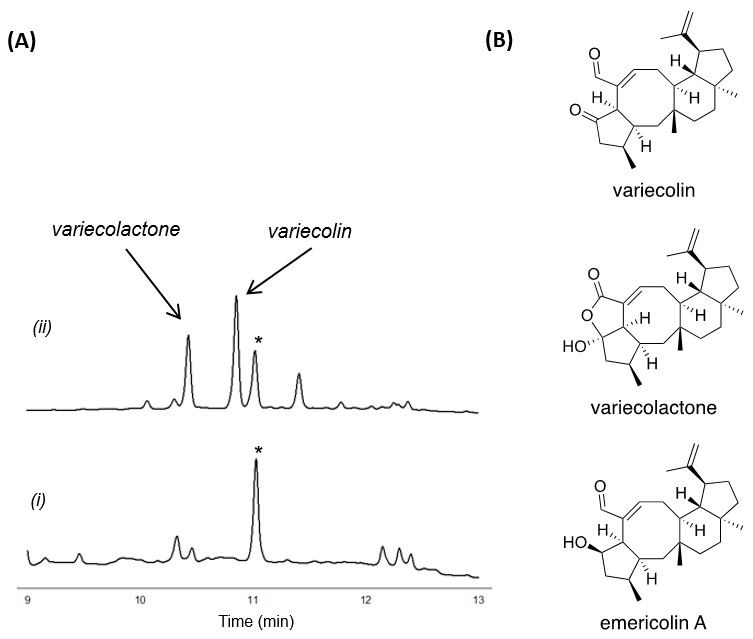
HPLC was used to confirm the presence of variecolin in the metabolites. The result indicated successful synthesis of variecolin, and further indicated that two other variecolin analogues, varicolactone and emericolin A, were also present in the metabolites; the peak of emericolin A overlapped with the peak of an endogenous metabolite derived from the A. oryzae strain. Their structures were further validated by NMR analysis as well as their specific rotation values, and the NMR data of the three compounds were in good agreement with the reported data [2][3][4].

Figure 2. (A) HPLC chromatograms of the metabolites from the A. oryzae transformants with (i) TS and (ii) TS and P450. Asterisk indicates an endogenous metabolite that co-elutes with emericolin A. The chromatograms were monitored at 215 nm. (B) Structures of the metabolites.
Learn
Cycle 2 - Biosynthesis of variecolin analogues
Transformation based on the random integration of target DNA requires a large amount of DNA and the screening of many transformants to find the most appropriate one. Because the introduction of individual P450 monooxygenase genes would be conducted in the next step, we employed a recently developed CRISPR-Cas9-mediated transformation method, coupled with homologous recombination, to facilitate the process.
Cycle 2 - Biosynthesis of variecolin analogues
Design - Selection of P450 homologues using cblaster

Because many fungal sesterterpenoid pathways adopt one or more P450 monooxygenases as a tailoring enzyme, we predicted that co-expression of a TS gene responsible for the synthesis of the variecolin scaffold and a homologue of variecolin synthase could yield a new variecolin analogue. Therefore, a number of fungal genomes containing P450 genes that are potentially associated with di-/sesterterpenoid pathways were selected in silico using the cblaster program [5]. A total of 17 cytochrome P450 genes were identified and selected for synthesis of novel variecolin analogues.

Figure 3. Part of the result of cblaster search.
Build - P450 gene transformation
P450 genes were PCR amplified from genomic DNA of different fungi and separately ligated to pTAex3-HR (a vector that contains the argB prototrophic gene marker) to generate a series of pTAex3-HR-P450 plasmids. The pTAex3-HR-P450 plasmids were separately transformed into the A. oryzae/TS strain by PEG-mediated protoplast transformation with the help of the CRISPR-Cas9 system.
CRISPR-Cas9 genome editing[6]
The essential components of CRISPR-Cas9 genome editing include guide RNA and Cas9. The guide RNA is designed to target the sC locus of the A. oryzaeNSAR1 strain. Guide RNA and Cas9 were added to the protoplast, causing the DNA double strand to break at the expected site. The double strand break can be repaired by homologous recombination using the pTAex3-HR-P450 plasmid. Because pTAex3-HR harbors the argB gene, and the resultant A. oryzae transformants become arginine prototroph, it can be selected on the transformation medium which lacks arginine. The introduction of the target P450 was verified by diagnostic PCR using a gene-specific primer, and an additional primer that attaches outside the homology arm used for the transformation.
Test - Analysis of the possible variecolin analogues

HPLC, LC-MS, and GC-MS analyses were performed to examine whether there were variecolin analogues in the A. oryzae transformants. Analysis results indicated that four of the tested P450s successfully converted the hydrocarbon product into oxidized metabolite(s). Although two of the four P450s exhibited the identical metabolite profile to that of the P450 from A. aculeatus, the other two afforded new metabolites, which were not present in the variecolin pathway. These metabolites were isolated via a series of chromatographic techniques, including flash chromatography and semipreparative HPLC. NMR and MS analyses were performed to determine the structures of the variecolin analogues. A total of three new variecolin analogues (1, 2, and 3) were successfully obtained.

Figure 4. HPLC chromatograms of the metabolites (1, 2, and 3)from the A. oryzae transformants with the artificial sesterterpenoid pathways.1: compound 313, 2: compound 161, and 3: compound 163.
Learn



The antibacterial and anticancer activities of the three new variecolin analogues were discovered and compared with the biological activities of variecolin. The result shows the potentially therapeutic properties of variecolin analogues; their other biological activities can be tested in the future.
Anti-bacterial properties
The in vitro antimicrobial activity of variecolin and variecolin analogues were tested with several microbial species, including Staphylococcus epidermidis, Staphylococcus aureus, Bacillus cereus, Enterococcus faecalis, and Escherichia coli. One of the variecolin analogues exhibited a minimum inhibitory concentration (MIC70) of 16 μg/ml on Staphylococcus aureus, compared to 7 μg/ml of variecolin.
Table 1. Anti-bacterial activities of purified secondary metabolites

Anticancer properties
The in vitro anticancer activity of variecolin and variecolin analogues was tested utilizing breast cancer cell MCF-7. The half maximal inhibitory concentration (IC50) of variecolin was 1.4±0.3 μM μM, and the new variecolin analogues exhibited anticancer properties with IC50 values of 8±3 μM, 31±9 μM, and 146±82(Table 2), respectively.
The inhibition of cell viability assay of breast cancer MCF-7 has shown that variecolin is the most efficient anticancer compound among all six compounds with IC50 value 1.4±0.3 μg/ml (Table 2). The new vairecolin analogue 313 showed inhibition of cell viability with IC50 value 8±3 μg/ml, which is stronger than that of the existing variecolactone with IC50 value 22±8 μg/ml. The new variecolin analogue 161 and 163 also showed anti-cancer properties against the breast cancer MCF-7 but weaker than that of variecolin and the new variecolin analogue 313. The low IC50 values of variecolin and the new variecolin analogue 313 indicate that they might be effective drugs for treating cancer and are worth further researching.
Table 2. Anticancer activities of purified secondary metabolites


Figure 5. %cell viability versus the concentration of purified metabolites
References
1. Jin, F. J., Maruyama, J. I., Juvvadi, P. R., Arioka, M., & Kitamoto, K. (2004). Development of a novel quadruple auxotrophic host transformation system by argB gene disruption using adeA gene and exploiting adenine auxotrophy in Aspergillus oryzae. FEMS Microbiology Letters, 239(1), 79-85.
2. Yoganathan, K., Rossant, C., Glover, R. P., Cao, S., Vittal, J. J., Ng, S., Huang, Y., Buss, A. D., & Butler, M. S. (2004). Inhibition of the human chemokine receptor CCR5 by variecolin and variecolol and isolation of four new variecolin analogues, emericolins A−D, from Emericella aurantiobrunnea. Journal of Natural Products, 67(10), 1681-1684.
3. Takahashi, H., Hosoe, T., Nozawa, K., & Kawai, K. I. (1999). Two new sesterterpenes from the ascomycetous fungus Emericella purpurea. Journal of Natural Products, 62(12), 1712-1713.
4. Yodsing, N., Lekphrom, R., Sangsopha, W., Aimi, T., & Boonlue, S. (2018). Secondary metabolites and their biological activity from Aspergillus aculeatus KKU-CT2. Current Microbiology, 75(5), 513-518.
5. Gilchrist, C. L., Booth, T. J., van Wersch, B., van Grieken, L., Medema, M. H., & Chooi, Y. H. (2021). cblaster: a remote search tool for rapid identification and visualization of homologous gene clusters. Bioinformatics Advances, 1(1), vbab016.
6. Wei, X., Matsuyama, T., Sato, H., Yan, D., Chan, P. M., Miyamoto, K., Uchiyama, M., & Matsuda, Y. (2021). Molecular and computational bases for spirofuranone formation in setosusin biosynthesis. Journal of the American Chemical Society, in press.
Copyright © 2021 - All Rights Reserved - VarieCure
Tech Supported by CS Dept, CityU


